BrainVoyager v23.0
Creation of High-Resolution VTCs
In most cases the voxel resolution of functional data is not below 1 mm and can, thus, be handled by the normal processing pipeline in BrainVoyager with 1 mm VMR data sets in native, MNI, AC-PC or Talairach space. When using sub-millimeter functional resolution, e.g. on ultra high-field scanners (7 Tesla and beyond), some special treatment is required in order to create sub-milimeter volume time course (VTC) data in AC-PC, MNI or Talairach space. Since sub-millimeter data is usually analyzed individually, it might be best to create VTCs in FMR-STC space, which avoids resampling to a normalized space. In this case the information in this topic is not needed and one should proceed with topic VTC Data in FMR Space..
The sub-millimeter functional data may be obtained from whole-brain scans but it is often acquired from small slabs; for the latter case, improved bounding box specification operations are available to avoid huge transformed datasets filled mainly with "empty" voxel time courses. Since functional data can not have a higher resolution than the "hosting" anatomical VMR data set (in the current framework of BrainVoyager), the easiest solution to handle sub-millimeter VTCs is to use anatomical data with sub-millimeter resolution. Note that it is not strictly necessary to measure the anatomical data with sub-millimeter resolution since it can be upscaled easily to any desired (iso-voxel) resolution. With the VMR data set in the right spatial resolution, the functional data can be created in the same resolution by using value "1" for the relative functional resolution value when creating VTC files (integral values 2, 3 and higher will produce functional data with lower resolution relative to the VMR as is often used with standard fMRI data). Since BrainVoyager QX 2.4 extensions to existing tools and file formats have been implemented that make it almost as easy to work with sub-millimeter anatomical and functional data as with standard resolution data in native, AC-PC, MNI and Talairach space. Further improvements in BrainVoyager QX 2.8 support specification, processing and visualization of Talairach space with any (sub-millimeter) spatial resolution. The steps below describe in detail how sub-millimeter VTCs can be created using data from a 7 Tesla scanner (courtesy of Federico de Martino, Maastricht University and Center for Magnetic Resonance Research, CMRR, Minneapolis, USA).
Creation of High-Resolution Spatial Transformation Files
Since BrainVoyager QX 2.8 it is possible and recommended to directly specify native, ACPC and Talairach transformations on the desired (sub-millimeter) resolution VMR data set that matches the resolution of the targeted sub-millimeter VTC resolution. A scaling to 1 mm VMR data for specifying ACPC transformation (.TRF) and Talairach landmark (.TAL) files (which was necessary in BrainVoyager QX 2.6), is no longer necessary since BrainVoyager now supports ACPC and Talairach space for any sub-millimeter voxel resolution. Also the advanced segmentation tools now process sub-millimeter ACPC/Talairach space data by rescaling the size of Talairach masks.
In order to run FMR-VMR alignment, ACPC transformation and Talairach warping, a high-resolution iso-voxel VMR data set is needed that matches the resolution of the desired sub-millimeter functional data since VTC files are created in the resolution of the "hosting" VMR. To ensure that this is indeed the case, the value of the functional-to-anatomical resolution must be set to 1x1x1 in the Create VTC Options dialog.
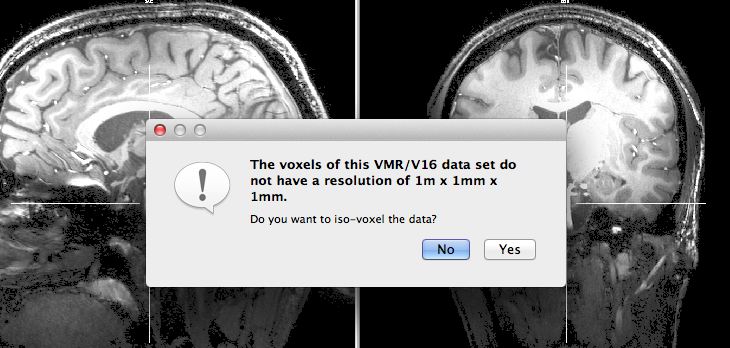
In the example data used to illustrate the procedure, functional data will be created with a resolution of 0.5 / 0.3 mm. The intra-session measured 3D anatomical data has a resolution of 0.3 mm. After creation of the VMR data set, the program asks whether the data should be iso-voxeled to a resolution of 1 x 1 x 1 mm (see snapshot above). Since we want to keep the native resolution, click the No button. Since the used 7 Tesla anatomical data set exhibits substantial spatial intensity inhomogeneities, they should be corrected first, i.e. before FMR-VMR alignment or cortex segmentation. In case one wants to resample the data at a later stage (to 1 mm or sub-millimeter iso voxels), the Iso-Voxel Transformation dialog can be invoked from the Spatial Transf tab of the 3D Volume Tools dialog.
VTC Creation
It is recommended to perform FMR-VMR alignment and ACPC / Talairach transformation at the native voxel resolution. While the automatic Talairach transformation is not yet available for sub-millimeter VMRs, landmarks for ACPC and Talairach transformation can be specified manually on sub-millimeter VMRs. Note that the partial or full Talairach bounding box will be visualized correctly at any voxel resolution (since BVQX 2.8). It is thus advised to proceed with the creation of ACPC and Talairach transformation (if desired) in the same way as described in the topic Manual ACPC and Talairach Transformation with the only difference that the high-resolution VMR is used instead of a 1 mm 3D anatomical data set. Also the FMR-VMR alignment files should be performed in the same way as described in topic Coregistration of Functional and Anatomical Data Sets selecting the high-resolution anatomical data set as the target VMR. With the obtained initial, fine-tuning, ACPC and Talairach transformation files, the high-resolution VTC data can then be created using the VTC File Creation dialog. Make sure that the dialog is called from the high-resolution VMR data set. If the functional-to-anatomical resolution is set to 1x1x1, the resulting VTC data will have the same resolution as the "hosting" VMR data set.
VTC Creation - An Alternative Approach
The approach described above was not available in older versions of BrainVoyager (prior to version 2.8). An alternative approach was described that uses resampled 1 mm anatomical data sets for ACPC and Talariach transformation instead of the native sub-millimeter resolution data. Although no longer recommended, this approach is described here in case it is needed (e.g. when using version 2.6 of BrainVoyager QX). In this approach, the high-resolution VMR data set is downsampled to a 1 x 1 x 1 mm VMR data set for FMR-VMR alignment and (automatic) Talairach transformation. This is best achieved by running the iso-voxelation tool with default settings, i.e. only the OK button needs to be clicked in the Iso-Voxel Transformation dialog. The program suggests as a file name the previous one with the new substring "_ISO"; in order to recognize that this data set is a 1 mm version, the trailing substring is changed to "_ISO1mm" for the example data set.
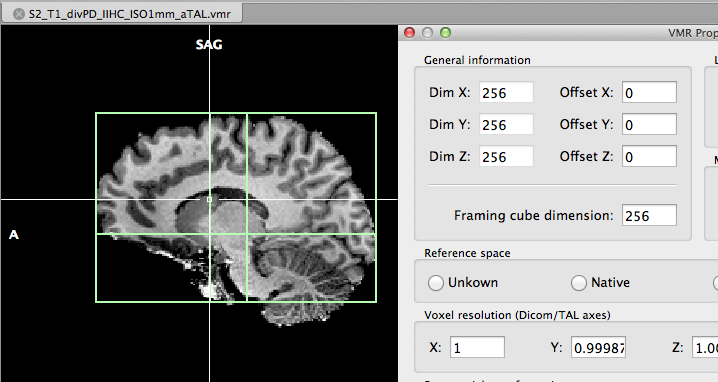
In order to transform the 1 mm VMR into ACPC and Talairach space, the automatic Talairach transformation can be started by clicking the Auto-ACPC-TAL button in the Talairach tab of the 3D Volume Tools dialog. The resulting "_aTAL" file is shown in the snapshot above. In case that the automatic transformation is not producing acceptable results, the interactive Talairach transformation can be used.
The transformation and TAL files created during Talairach transformation store information about the resolution and framing space used. This is important since this supports application of the transformations during VTC creation for other resolutions by performing proper scaling of the functional data. The "_aACPC.trf" file, for example, ends with these lines
ACPCVMRFramingCube: 256 ACPCVMRVoxelRes: 1
that inform BrainVoyager about the voxel resolution and (framing) dimensions used when this transformation file was created. In case that a 0.5 mm data set would be used, the "ACPCVMRFramingCube" would be "512" and the "ACPCVMRVoxelRes" entry would be "0.5". Similarly as for spatial transformation (.TRF) files, also the Talairach (.TAL) landmark files keep relevant resolution information. The created "_aACPC.tal" file, for example, ends with these lines:
TALVMRFramingCube: 256 TALVMRVoxelRes: 1
In case that this Talairach file is used with a 0.3 or 0.5 mm VMR data set, the defined coordinate values for AC, PC etc. will be scaled accordingly to create functional (VTC) data in the appropriate space. The final files that are needed to create VTC data are the FMR-VMR alignment files. In the context of the used example data, a functional project has been created from intra-session data of a conducted auditory experiment. A snapshot of the preprocessed FMR project is shown below. The resolution of the functional data is 1.5 x 1.5 x 1.5 mm, i.e. it is not a sub-millimeter data set but for demonstration purposes we will create a sub-millimeter VTC data set in Talairach space.
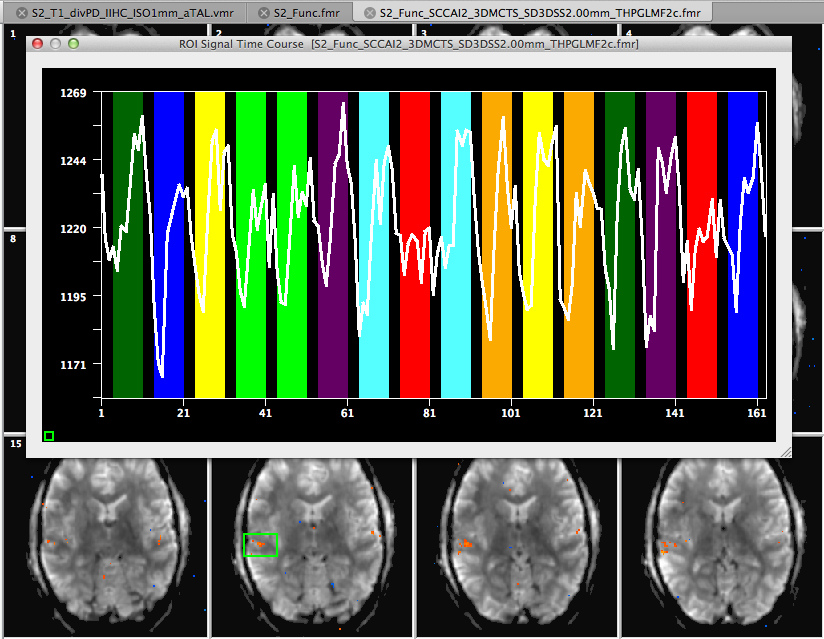
The FMR-VMR coregistration is performed in the usual way, i.e. by executing an initial and a fine-tuning alignment step from the 1mm VMR in native space and with the functional data (FMR) as input.
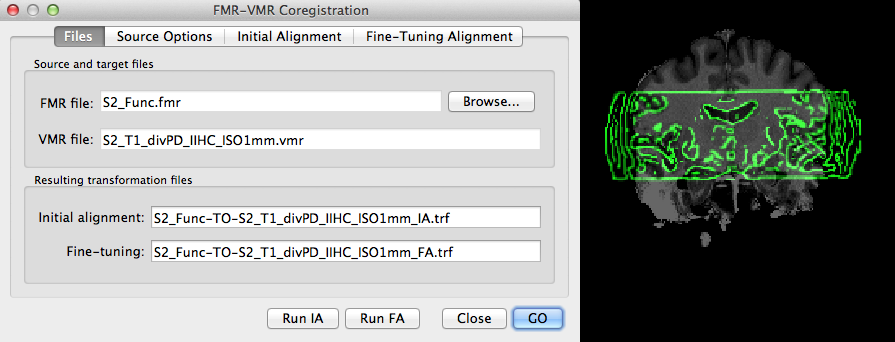
The snapshot above shows the used input files and the produced output files in the FMR-VMR Coregistration dialog (left side) and a coronal slice showing the obtained high-quality coregistration (right side). The obtained "_IA.trf" and "_FA.trf" files also keep information about the resolution of the hosting VMR file used during alignment:
ToVMRFramingCube: 256 ToVMRVoxelRes: 1
Application of Transformation Files with Sub-Millimeter VMR
With the resolution entries in all spatial transformation files, the program knows what extra adjustments to perform (e.g. scalings) when creating VTC files in a space defined by a given VMR file, e.g. with 256 or 512 framing dimensions. If, for example, the alignment would be done with a (scaled) 0.5 mm VMR (512 framing cube), the program would create a proper transformation matrix to create a VTC file in the 512 framing space. Since there are 1 mm and 0.3 mm VMRs in our example, VTCs can be created in one of these spaces depending on the chosen native space VMR. Since a VMR can be scaled to any other space (e.g. 0.5 mm), this implies that VTCs can be created in any desired resolution. In the context of our example data, we will create a VTC file using the high-resolution VMR file with a resolution of 0.3 mm and framing dimensions of 768.
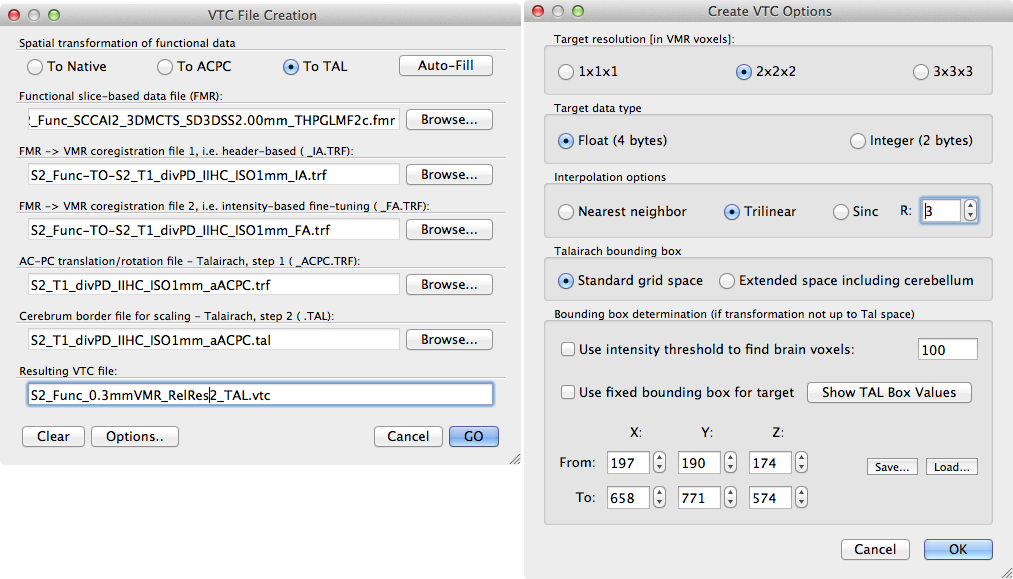
The snapshot above shows the standard VTC File Creation dialog (left side) that was filled automaticlly after selectiong the functional .fmr file as input followed by a click on the Auto-Fill button. The resulting VTC file name has been edited to reflect that the used VMR has 0.3mm resolution. Note that a VTC file is created with a relative resolution with respect to the VMR. The Create VTC Options dialog can be used to specify the relative resolution in the Target resolution [in VMR voxles] field. If the 1 x 1 x 1 option is chosen, the VTC will have the same resolution as the "hosting" VMR (0.3 mm in our example). Since this will lead to a huge VTC, the 2 x 2 x 2 option has been chosen for this example that will create an effective resolution of 2 x 0.3 mm = 0.6 mm in each dimension. This resolution is still highly oversampling the true resolution (1.5 mm) of the functional data; this is not recommended but only used here for demonstration purposes. Even with a resolution of 0.6 x 0.6 x 0.6 mm, the resulting VTC file has a size of 8.7 GB on disk! In order to visualize and analyze the 0.6 mm VTC data in Talairach space, the 1 mm Talairach ("_aTAL.vmr") file needs to be also available in 0.3 mm VMR resolution. This can be achieved either by applying the ACPC .trf file to the 0.3 mm VMR in native resolution followed by application of the .TAL file or by simply upscaling the 1 mm Talairach VMR file. The latter can be performed easily using the same Iso-Voxel Transformation dialog as used above.
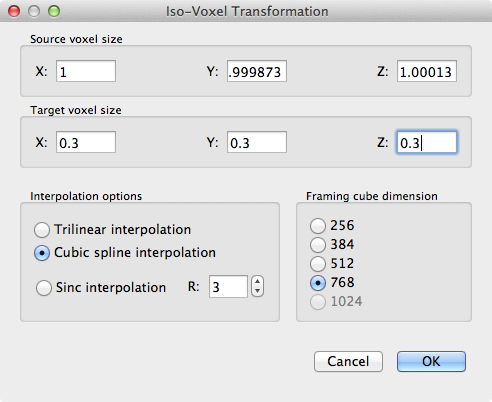
The snapshot above shows this dialog with a target voxel size of 0.3 x 0.3 x 0.3 mm; note that the framing cube in the Framing cube dimension field needs to be set to 768 so that the final VMR is large enough to hold the upscaled data set. The resulting 0.3 mm VMR can then be used to link the 0.6 mm VTC file. The snapshot below shows an overlay of a statistical GLM map calculated from the 0.6 mm VTC data superimposed on the 0.3 mm VMR data set.
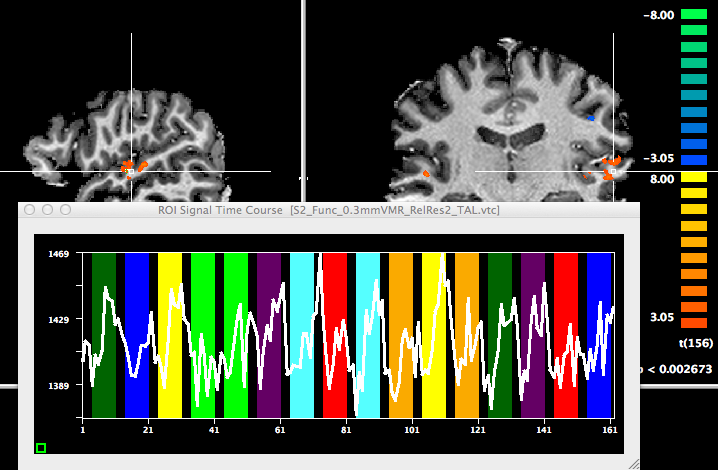
Limiting Sub-Millimeter VTC to Specified Bounding Box
In order to avoid huge VTCs when only measuring (or being interested in) part of the brain, it is advisable to create VTC data sets only within a bounding box covering the interested sub-space of the brain. The context menu in the VMR sub-views has an entry called Set Bounding Box that can be used to quickly define a bounding box around a voxel selected by the voxel selection cross (see snapshot below).
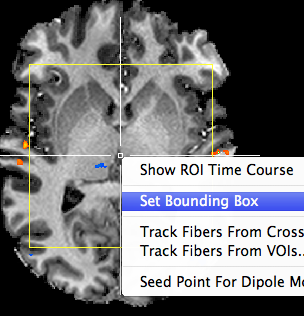
The size of the bounding box can be fine-adjusted using the spin boxes in the Bounding box field of the Segmentation tab of the 3D VolumeTools dialog. Any changes in the respective spin boxes will be visualized immediately in the VMR sub-views in case that the Show option in the Bounding box field is turned on (default). In order to quickly adjust the borders of the box, one can also use the keyboard up/down keys that advance the selected bounding box border (active spin box) in steps of 10 units.
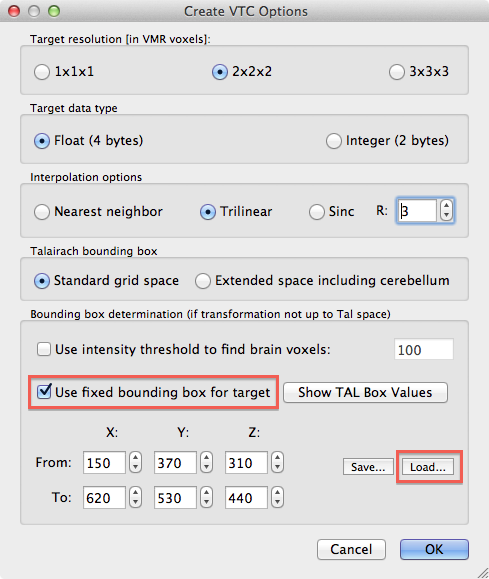
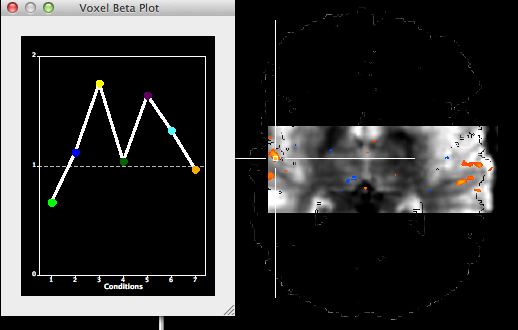
If the bounding box specification has been completed, it may be saved to disk using the Save button in the Bounding box field as a .bbx file. Most importantly, the saved .bbx file can be loaded in in the Create VTC Options dialog using the Load button in the Bounding box determination field to restrict the resulting VTC file to the desired sub-space. Note also that using the same bounding box from a stored .bbx file allows to apply the same sub-space for each subject of a group study within TAL space. For the example data, a bounding box covering a slab containing the left and right auditory cortex has been defined and then loaded for creation of the sub-millimeter VTC sub-space data (see snapshot above). When loading a .bbx file, the Use fixed bounding box for target option is turned on automatically but if the bounding box values are entered manually, this option must be turned on explicitly. In our example case, the resulting sub-space VTC data has a size of only 723 MB on disk as compared to the 8.7 GB for the full Talairach space file. The snapshot above shows the selected slab in Talairach space that has been used to create the 0.6 mm VTC file.
Note. The definition of the bounding box is invariant with respect to using a VMR with full framing cube dimensions or when using a VMR with minimized dimensions (as long as the same framing cube size is used) since the program adjusts the bounding box definition with the offset values of the current VMR when saving the .bbx file to disk.
Copyright © 2023 Rainer Goebel. All rights reserved.