BrainVoyager v23.0
Creating Sulcal Depth Maps
This topic describes how one can create and visualize sulcal depth volume and surface maps that provide information about the depth of voxels/vertices within sulci. While this information is interesting in itself for analysis, it provides visualizations of the macroanatomy of the folded cortex for inflated meshes and flat maps that can be used instead of the curvature-based standard visualization.
Preparing VMR for Sulcal Depth Distance Calculation
In order to calculate sulcus depth maps, we need to prepare an appropriate VMR data set that contains two segmentations identified by two intensity values ("colors"). The first segmentation is used to specify the starting region for the distance calculation. For sulcal depth measurements this needs to be (roughly) the outer contours of the brain without extensions into the sulci (convex hull). Calculating a distance transform from this first segmentation will provide a shortest distance value from the start region at each voxel in the volume. In order to stop/limit the distance transformation at banks of sulci, a second segmentation is used. For the demonstration below this second segmentation is the white matter / gray matter boundary but one can also use a mid-GM volume (after advanced segmentation) or a GM/CSF boundary.
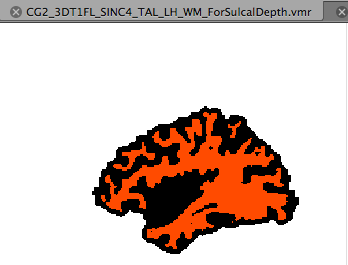
Both segmentations can be prepared from one segmentation file, e.g. the “WM” VMR of one hemisphere resulting from the standard cortex segmentation tools (a mid-GM or pial segmentation could also be used but the pial volume might not have “open” sulci everywhere and it is thus recommended to work with the WM volume). Simply load the “WM” segmentation of the left or right hemisphere of the desired data set. Then open the 3D Volume Tools Options dialog from the Segmentation tab of the 3D Volume Tools dialog. Switch to the Operations tab where you will find the Distance transform group box. To start the sulcal depth preparation step, click the Prepare Sulcal Depth button. The program will now apply morphological operations (dilation and erosions) in order to close the sulci producing a segmentation roughly at the outer brain boundary; the segmentation is then inverted so that the voxels within the brain are black (intensity 0) and the space outside the brain appears white (intensity 255); finally the original WM segmentation is put back into the resulting volume using a red/orange color (intensity 226). After about a minute, you should see the resulting VMR in the BrainVoyager workspace that is auomatically saved to disk using the original file name with the extension ”_ForSulcalDepth.vmr“ (see snapshot above).
Creation of sulcal depth volume maps
With the “_ForSulcalDepth.vmr” VMR created in the previous step, a distance transform can now be calculated resulting in values at each voxel that measure the shortest distance from that voxel to the white (background) tissue; for the prepared data, this will result in a volume map containing sulcal depth values at each voxel in the “black” tissue.
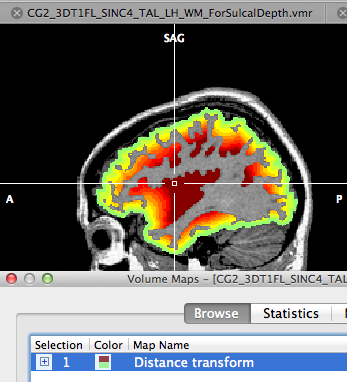
Make sure that the sulcal depth VMR prepared in the previous section is the current VMR. Then go to the same tab (Operations) as before in the 3D Volume Tools dialog and click the Distance Map button. The launched distance transformation routine expects the colors detailed above as input, i.e. it localizes the starting boundary of the distance calculation at the transition from white to black voxels and it calculates the shortest distance to each voxel in the black space between the white and red (i.e. WM) segmentations. After a short moment the calculated distance volume map will appear in the current VMR window overlayed on the prepared VMR. The distance map is also automatically saved to disk under the prepared VMR name, however, with an appropriate extension of “_DistanceMap.vmp”. To better visualize the obtained map, it is recommended to open the original (or IIHC corrected) VMR data set and to load the stored map on this VMR (see snapshot above). If you navigate through the volume you will find color-coded voxels superimposed on gray matter voxels that code the depth of a voxel with respect to the outer brain boundary. Shallow depth values are coded with green colors, depth values of 5 - 10 mm are color coded with yellow/orange colors and very deep values (15 and higher) are color coded with dark red values. Of course you can change this mapping using the standard upper range (threshold) value and you can also change the used color look-up table.
Creation of sulcal depth surface maps
For visualization and further analysis, the obtained sulcal depth volume map can be mapped on to a cortex mesh creating a sulcal depth surface map. With the current VMR visualizing the volume map, open a cortex mesh on which you want to project the volume map. It is recommended to use the standard “RECOSM” surface of the corresponding hemisphere (LH or RH) for this purpose but also a mid-GM mesh (available when using the advanced segmentation tools) or pial cortex mesh can be used. A mesh generated for cortex-based alignment ("SPH") can also be used allowing to create group averages of depth maps among other analyses. To create the surface map from the active volume map, enter the Surface Maps dialog and click the Create SMP button. This will present the Depth Integration dialog that can be used to either accept or modify how the mesh should sample the prepared volume map. Since we want to sample along the mesh without integrating in depth, we could use the option Sampling volume data exactly along mesh vertices. Since the RECOSM mesh runs, however, pretty close to the WM/GM boundary, some modificatins are suggested (see snapshot below).
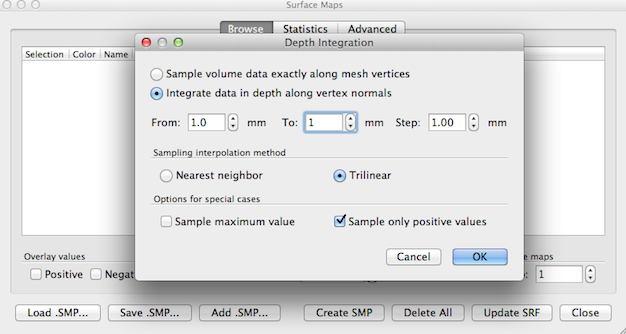
First, the default option Integrate data in depth along vertex normals should be kept but with changes to the From and To values that should both be set to 1.0. This in effect will sample along the vertices (no integration in depth) but with a displacement of 1 mm along a vertex normal, i.e. 1 mm into grey matter; this will produce better results at the vertices close to the outer brain boundary. It is also recommended to turn on the special option Sample only positive values; the reason is that the white matter voxels in the volume map (where the distance transform had been stopped) have gotton values of ”–1“ and these values may become (partially) included in the sampled depth values extracted from the volume map at a given mesh vertex, especially when the Trilinear interpolation option is used. After accepting the (modified) settings with a click on the OK button, the surface map will be created and visualized on the mesh (see snapshot below). There might be a few vertices without a map value (”holes“) that can be ”closed“ by spatial mesh smoothing. To do this, go to the Advanced tab of the Surface Maps dialog and click the Smooth SMP button in the Smooth selected map field (since one smoothing step is usually enough, you may want to reduce the number of smoothing iterations from 5 to 1 in order to produce minimal changes of the surface map).
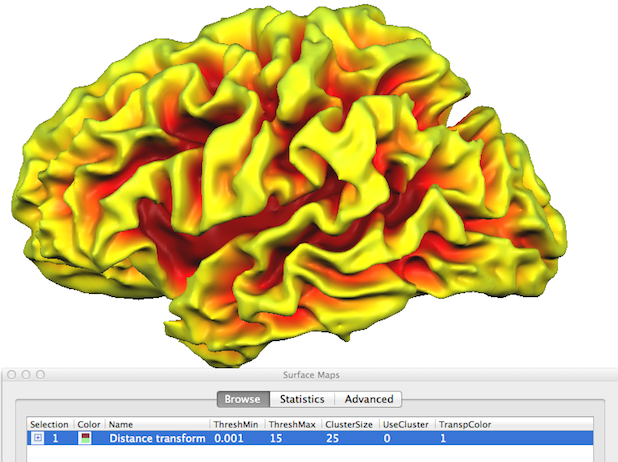
The created sulcal depth surface map can be explored easily by using CTRL/CMD mouse clicking on an interesting region; this will show the "sampled" surface map value (i.e. depth value) in the status bar. In order to better see and select desired regions, it may be helpful to inflate the mesh (see snapshot below).
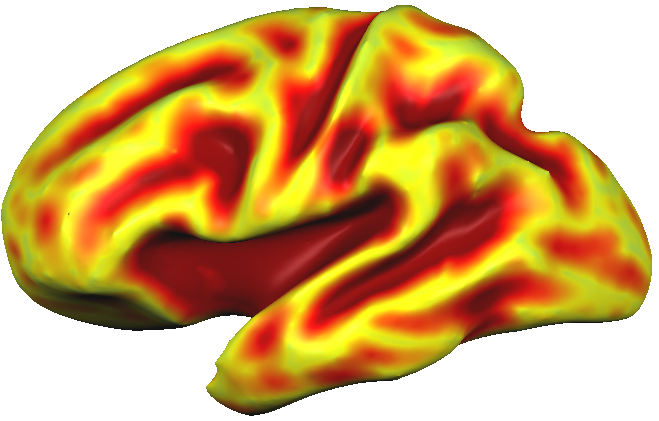
By chosing a grey scale color look-up table, a sulcal depth map may also be an alternative to highlight macroanatomy (gyri and sulci) but with less detail as with curvature-based visualizations. In the snapshot below, the "grey_sulcaldepth.olt" LUT was used and the upper threshold value was set to 9.0.
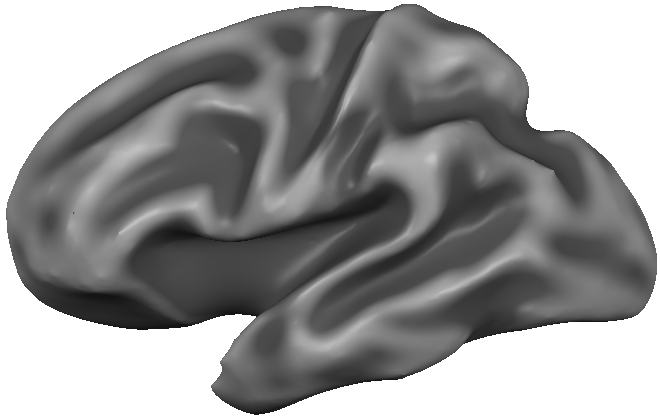
Using POIs, detailed regional information may be extracted from patches or line paths such as POIs defined along the fundus of sulci (see snapshot). This can be, for example, helpful to compare the average (or peak) depth value across corresponding sulci in the left and right hemisphere.
Copyright © 2023 Rainer Goebel. All rights reserved.