BrainVoyager v23.0
Creation of Topologically Correct Cortex Segmentations
The result of the automatic white / grey segmentation (see previous steps) can be further improved by removing topological errors called "bridges" or "handles" using a "bridge removal" algorithm (Kriegeskorte & Goebel, 2001).

The files "[name]_TAL_LH_WM.vmr" and "[name]_TAL_RH_WM.vmr" produced in the previous step can be directly used to reconstruct its boundaries producing a mesh representation of the cortical sheet of each hemisphere. The quality of the resulting polygon meshes is well suited to visualize the folded cortical sheet in the surface module. It is also well suited to calculate cortex masks to be used for "cortex-based statistics" by limiting statistical analyses to cortical voxels.
The created meshes are, however, in most cases not appropriate for further morphing procedures like inflation, flattening and creation of spherical representations because they are not completely free of topological errors called "handles" or "bridges". These topological errors are often the result of connections across opposite banks of sulci or of holes within sulcal banks. These bridges can be removed by manual editiing as described in section xx. It is also possible to remove these errors automatically using the "bridge removal" tool (for details, see Kriegeskorte & Goebel, 2001). To use this tool, you have to enable the Remove bridges LH and/or the Remove bridges RH option in the Automatic Cortex Segmentation and Reconstruction dialog (turned off as default). The resulting topologically correct data sets will be saved to disk as "[name]_LH_WM_BL2.vmr" and "[name]_RH_WM_BL2.vmr" and are used for surface reconstruction of the cortical sheet.
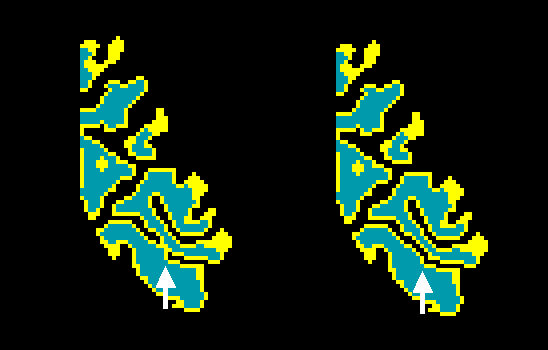
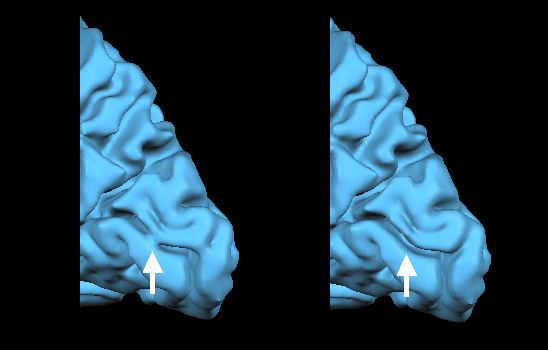
The figure above shows part of a topological error, a "bridge", in the calcarine sulcus and the same region after the automatic bridge removal algorithm has been applied. The figure below shows the same region in the corresponding polygon mesh representation in the 3D Viewer window after surface reconstruction.
While the bridge removal tool guarantees that the resulting segmentation is topologically correct, it can not correct certain types of errors, such as partial inclusion of grey matter regions. These additional errors occur in parahippocampal regions and in the amygdala and should be manually corrected before or after removal. In some cases, also the ventricles are not properly filled. These kind of errors can not be corrected by the bridge removal algorithm. Note, however, that you can rerun the bridge removal algorithm after manual editing the segmented volumes from the Segmentation Options dialog (see snapshot below), which you can launch by clicking the Options button in the Segmentation tab of the 3D Volume Tools dialog.
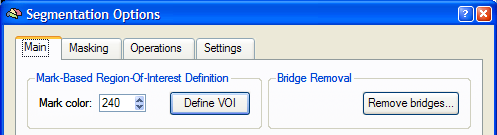
After clicking the Remove Bridges button in the Main tab of the Segmentation Options dialog, the Bridge Removal dialog appears (see below). Before starting the bridge removal process, you should specify the original (unsegmented) data set in the Source file field using the corresponding Browse button. The Presegmented file field shows the name of the currently loaded VMR file, which will be most likely one of the two segmented hemispheres. Since the bridge removal algorithm decides whether to apply cuts or fills to correct topological errors using intensity information, it is important to provide information about the threshold between grey and white matter. This can be easily done by clicking the Estimate button, which will fill the fields Grey matter, White matter and Threshold after inspection of the intensity histogram of labelled white and neighboring grey matter voxels. If the Show histogram option is checked (default), a plot of the intensity histogram is also shown as additional information.
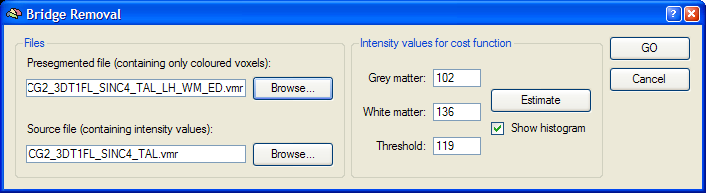
To run the bridge removal algorithm for the current hemisphere, click the GO button. The process will last about five minutes on current computer hardware.
References
Kriegeskorte, N. & Goebel, R. (2001). An efficient algorithm for topologically correct segmentation of the cortical sheet in anatomical MR volumes. Neuroimage, 14, 329-346.
Copyright © 2023 Rainer Goebel. All rights reserved.